In January 2020, the first complete severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) genome sequence was deposited to GenBank. Since then, the amount of new genome sequences has increased rapidly at GenBank and GISAID. This has laid the foundations for analyzing the virulence, antigenicity, pathogenicity, and transmissibility of SARS-CoV-2 mutations.
Structural and non-structural proteins of SARS-CoV-2
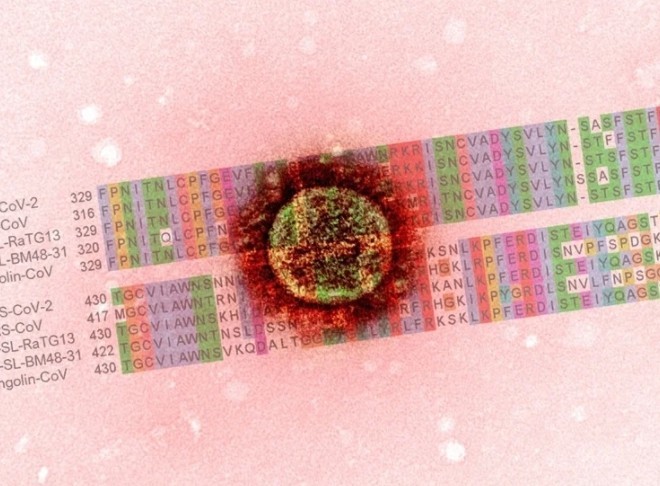
SARS-CoV-2 is a positive-sense single-stranded RNA virus that encodes 29 structural and non-structural proteins (NSPs) using 29,903 nucleotides. The structural proteins help with the formation of the viral particle, while the NSPs play a significant role in replicating viral RNA. SARS-CoV-2 has 4 structural proteins, namely, spike (S), membrane (M), envelope (E), and nucleocapsid (N) proteins.
Of these, the spike (S) protein has 1,273 residues of SARS-CoV-2 and has a critical role in viral infection. It is essential for interaction with host cell receptors and the fusion of the viral envelope with the host cell membrane to allow entry of the virus. It is also the site of most SARS-CoV-2 mutations. Hence scientists have been focusing their attention on the Spike protein to develop antibody-based drugs and vaccines.
Identifying other fast-growing mutations on the SARS-CoV-2 spike (S) protein
The new variants of SARS-CoV-2 from the UK, Brazil, and South Africa have garnered a lot of attention recently for their increased infectivity, possibly high pathogenicity, and potential threats to currently available vaccines and antibody-based therapies. However, it is not clear if there are more variants with higher infectivity that are being transmitted across the world.
Researchers from Michigan State University recently carried out a large-scale study with 506,768 SARS-CoV-2 genome isolates gathered from COVID-19 patients. Their objective was to identify other rapidly growing mutations on the SARS-CoV-2 spike (S) protein receptor-binding domain (RBD). The research is published in the journal Genomics.
Commonly observed mutations strengthen the binding between RBD and host ACE2 receptor
The findings reveal that all 100 of the most observed mutations strengthen the binding between the RBD and the angiotensin-converting enzyme 2 (ACE2) of the host, which indicates that SARS-CoV-2 evolves into more infectious variants. In particular, this enhancement of the binding between RBD and ACE2 is observed in fast-growing RBD mutations such as N439K, S477N, S477R, and N501T.
The researchers further found that the N501Y mutation found in the UK, Brazil, and South Africa variants may modestly weaken the binding between many known antibodies and the RBD. The E484K and K417N mutations found in the Brazilian and South African variants and the L452R and E484Q mutations found in the Indian variants of SARS-CoV-2 can disrupt the binding between the RBD and many antibodies. The L452R RBD mutation is also known to be a part of the California variant named B.1.427.
T478K mutation makes the Mexico strain the most infectious SARS-CoV-2 variant
The mutations most likely to have vaccine-escape capabilities include S494P, K417N, Q493L, F490S, R403K, F486L, L452R, E484K, K417T, E484Q, F490L, and A475S. Also, the T478K mutation seems to make the B.1.1.222 variant found in Mexico the most infectious variant. According to the authors, the RBD mutations that can simultaneously disrupt the existing antibodies (vaccine escape mutations) and make SARS-CoV-2 more infectious may pose a looming threat to the current set of approved vaccines.
The comprehensive genetic analysis and protein-protein binding study by the authors show that the SARS-CoV-2 genetic evolution on the RBD may be regulated by viral proofreading, host gene editing, natural selection, and random genetic drift. This gives rise to more infectious SARS-CoV-2 variants that may compromise existing antibody-based COVID-19 treatment strategies and vaccines.


